
Grounded in the liberal arts, the Morrissey College offers undergraduate and graduate programs in the humanities, social sciences, and natural sciences, preparing students to thrive in a broad range of careers while using their gifts to enhance the common good.
37
Majors
48
Minors
22
Academic Departments
4
Special Programs
BC's approach to liberal arts education challenges students to learn across disciplines, prepares them for a range of rewarding careers, and helps them discern who they want to be—and why.
The centerpiece of Jesuit education has always been a common curriculum that emphasizes the rigorous study of the defining works of the humanities, natural sciences, and social sciences.
25+
Graduate Programs
585+
Doctoral Students
190+
Master's Students
100%
Funding for Doctoral Students
Our community is made richer by the diversity of our students, faculty, and staff, whose unique perspectives and lived experiences contribute to the vibrancy of our campus and classrooms.
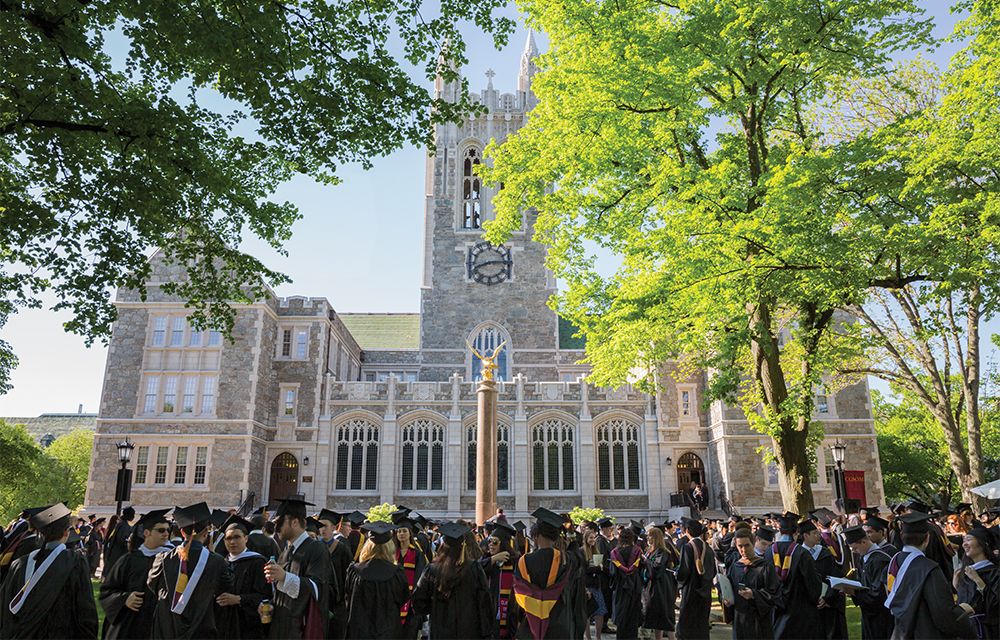















Boston College is at the forefront of scientific research and innovation, while also prioritizing a formative liberal arts education as the foundation for our R1 Research acumen. Across disciplines, collaborative teams of students and faculty are examining the complex problems of our contemporary world, proposing new solutions and new ways of thinking about religion and culture, science and technology, art and education, business and ethics.
To make a donation to the Morrissey College of Arts and Sciences, please visit the University's Giving website.